Page 46 - Read Online
P. 46
Page 413 Aydin et al. J Transl Genet Genom. 2025;9:406-26 https://dx.doi.org/10.20517/jtgg.2025.108
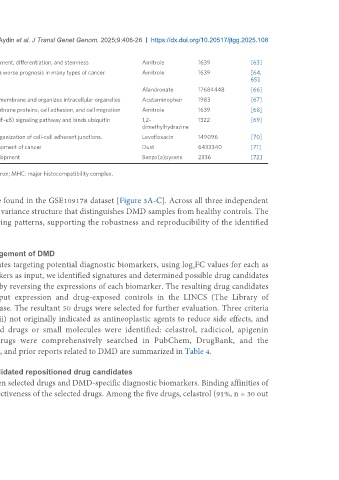
SOX4 Q06945 UP It is a crucial transcription factor for controlling progenitor development, differentiation, and stemness Amitrole 1639 [63]
SP1 P08047 TF It acts as a transcription factor, and its overexpression is linked to a worse prognosis in many types of cancer Amitrole 1639 [64,
65]
SPP1 P10451 UP It is involved in osteoclast binding to the mineralized bone matrix Alandronate 17684448 [66]
SPTAN1 Q13813 UP It functions as a pivotal scaffold protein that stabilizes the plasma membrane and organizes intracellular organelles Acetaminophen 1983 [67]
SPTBN1 Q01082 UP-DOWN It plays a role in determining cell shape and in regulating transmembrane proteins, cell adhesion, and cell migration Amitrole 1639 [68]
SQSTM1 Q13501 UP-DOWN Encoded protein controls the activation of the nuclear factor-κB (NF-κB) signaling pathway and binds ubiquitin 1,2- 1322 [69]
dimethylhydrazine
SSX2IP Q9Y2D8 UP It acts as a centrosome maturation factor and plays a role in the organization of cell-cell adherent junctions. Levofloxacin 149096 [70]
TP53 P04637 TF It is a tumor suppressor that plays a fundamental role in the development of cancer Dust 6433340 [71]
TWIST1 Q15672 UP-DOWN It regulates cell migration and proliferation during embryonic development Benzo(a)pyrene 2336 [72]
DMD: Duchenne muscular dystrophy; PPI: protein-protein interaction; NF-κB: nuclear factor-κB; IFN: interferon; MHC: major histocompatibility complex.
TWIST1 genes in the GSE38417 dataset; PLXND1, CD44, and GLS genes were found in the GSE109178 dataset [Figure 3A-C]. Across all three independent
datasets, PCA shows that the identified network biomarkers exhibit a coherent variance structure that distinguishes DMD samples from healthy controls. The
variable contribution maps further reveal gene-specific influences on clustering patterns, supporting the robustness and reproducibility of the identified
biomarker signatures.
Drug repositioning analysis revealed candidate therapeutics for the management of DMD
2
The L1000CDS search engine was used to identify repositioned drug candidates targeting potential diagnostic biomarkers, using log FC values for each as
2
signature inputs. Using the genes and FC values of potential diagnostic biomarkers as input, we identified signatures and determined possible drug candidates
that could reverse gene expression. A healthy state might have been achieved by reversing the expressions of each biomarker. The resulting drug candidates
were ranked regarding 1-cosα values, which indicate overlap between input expression and drug-exposed controls in the LINCS (The Library of
Integrated Network-Based Cellular Signatures, https://lincsproject.org/) database. The resultant 50 drugs were selected for further evaluation. Three criteria
were applied to prioritize repositioned drugs: (i) preferably FDA approved, (ii) not originally indicated as antineoplastic agents to reduce side effects, and
(iii) high 1-cosα values. Applying these criteria, five potential repositioned drugs or small molecules were identified: celastrol, radicicol, apigenin
triacetate, emetine dihydrochloride hydrate, and withaferin-A. These drugs were comprehensively searched in PubChem, DrugBank, and the
relevant literature. Their indications, mechanisms of action, approval statuses, and prior reports related to DMD are summarized in Table 4.
Molecular docking analysis indicated celastrol and emetine as in silico validated repositioned drug candidates
We performed molecular docking analyses to determine the interactions between selected drugs and DMD-specific diagnostic biomarkers. Binding affinities of
inhibitors of these biomarkers were used as positive controls to evaluate the effectiveness of the selected drugs. Among the five drugs, celastrol (91%, n = 30 out