Page 48 - Read Online
P. 48
Page 415 Aydin et al. J Transl Genet Genom. 2025;9:406-26 https://dx.doi.org/10.20517/jtgg.2025.108
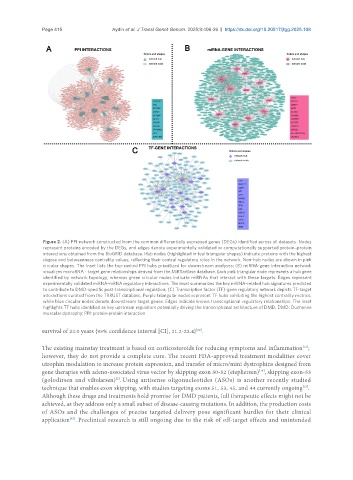
Figure 2. (A) PPI network constructed from the common differentially expressed genes (DEGs) identified across all datasets. Nodes
represent proteins encoded by the DEGs, and edges denote experimentally validated or computationally supported protein–protein
interactions obtained from the BioGRID database. Hub nodes (highlighted in teal triangular shapes) indicate proteins with the highest
degree and betweenness centrality values, reflecting their central regulatory roles in the network. Non-hub nodes are shown in pink
circular shapes. The inset lists the top-ranked PPI hubs prioritized for downstream analyses; (B) miRNA-gene interaction network
visualizes microRNA - target gene relationships derived from the MiRTarBase database. Each pink triangular node represents a hub gene
identified by network topology, whereas green circular nodes indicate miRNAs that interact with these targets. Edges represent
experimentally validated miRNA-mRNA regulatory interactions. The inset summarizes the key miRNA-related hub signatures predicted
to contribute to DMD-specific post-transcriptional regulation; (C) Transcription factor (TF)-gene regulatory network depicts TF-target
interactions curated from the TRRUST database. Purple triangular nodes represent TF hubs exhibiting the highest centrality metrics,
while blue circular nodes denote downstream target genes. Edges indicate known transcriptional regulatory relationships. The inset
highlights TF hubs identified as key upstream regulators potentially driving the transcriptional architecture of DMD. DMD: Duchenne
muscular dystrophy; PPI: protein-protein interaction.
survival of 22.0 years {95% confidence interval [CI], 21.2-22.4} .
[82]
The existing mainstay treatment is based on corticosteroids for reducing symptoms and inflammation ;
[12]
however, they do not provide a complete cure. The recent FDA-approved treatment modalities cover
utrophin modulation to increase protein expression, and transfer of micro/mini dystrophins designed from
[14]
gene therapies with adeno-associated virus vector by skipping exon 50-52 (etepliersen) , skipping exon-53
[2]
(golodirsen and viltolarsen) . Using antisense oligonucleotides (ASOs) is another recently studied
technique that enables exon skipping, with studies targeting exons 51, 53, 45, and 44 currently ongoing .
[13]
Although these drugs and treatments hold promise for DMD patients, full therapeutic effects might not be
achieved, as they address only a small subset of disease-causing mutations. In addition, the production costs
of ASOs and the challenges of precise targeted delivery pose significant hurdles for their clinical
application . Preclinical research is still ongoing due to the risk of off-target effects and unintended
[83]