Page 44 - Read Online
P. 44
Singh et al. Cancer Drug Resist. 2025;8:56 Page 9 of 20
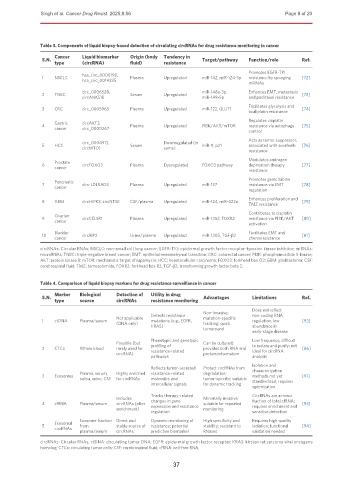
Table 3. Components of liquid biopsy-based detection of circulating circRNAs for drug resistance monitoring in cancer
Cancer Liquid biomarker Origin (body Tendency in
S.N. Target/pathway Function/role Ref.
type (circRNA) fluid) resistance
Promotes EGFR-TKI
hsa_circ_0000190,
1 NSCLC Plasma Upregulated miR-142, miR-124-3p resistance by sponging [72]
hsa_circ_0014235
miRNAs
circ_0006528, miR-148a-3p, Enhances EMT, metastasis
2 TNBC Serum Upregulated [73]
circANKS1B miR-149-5p and paclitaxel resistance
Facilitates glycolysis and
3 CRC circ_0005963 Plasma Upregulated miR-122, GLUT1 [74]
oxaliplatin resistance
Regulates cisplatin
Gastric circAKT3,
4 Plasma Upregulated PI3K/AKT/mTOR resistance via autophagy [75]
cancer circ_0000267
control
Acts as tumor suppressor;
circ_0004913, Downregulated (in
5 HCC Serum miR-9, p21 associated with sorafenib [76]
circMTO1 some)
resistance
Modulates androgen
Prostate
6 circFOXO3 Plasma Dysregulated FOXO3 pathway deprivation therapy [77]
cancer
resistance
Promotes gemcitabine
Pancreatic
7 circ-LDLRAD3 Plasma Upregulated miR-137 resistance via EMT [78]
cancer
regulation
Enhances proliferation and
8 GBM circHIPK3, circNT5E CSF/plasma Upregulated miR-124, miR-422a [79]
TMZ resistance
Contributes to cisplatin
Ovarian
9 circCELSR1 Plasma Upregulated miR-1252, FOXR2 resistance via PI3K/AKT [80]
cancer
activation
Bladder Facilitates EMT and
10 circRIP2 Urine/plasma Upregulated miR-1305, TGF-β2 [81]
cancer chemoresistance
circRNAs: Circular RNAs; NSCLC: non-small cell lung cancer; EGFR-TKI: epidermal growth factor receptor-tyrosine kinase inhibitor; miRNAs:
microRNAs; TNBC: triple-negative breast cancer; EMT: epithelial-mesenchymal transition; CRC: colorectal cancer; PI3K: phosphoinositide 3-kinase;
AKT: protein kinase B; mTOR: mechanistic target of rapamycin; HCC: hepatocellular carcinoma; FOXO3: forkhead box O3; GBM: glioblastoma; CSF:
cerebrospinal fluid; TMZ: temozolomide; FOXR2: forkhead box R2; TGF-β2: transforming growth factor beta 2.
Table 4. Comparison of liquid biopsy markers for drug resistance surveillance in cancer
Marker Biological Detection of Utility in drug
S.N. Advantages Limitations Ref.
type source circRNAs resistance monitoring
Does not reflect
Non-invasive;
Detects resistance non-coding RNA
Not applicable mutation-specific
1 ctDNA Plasma/serum mutations (e.g., EGFR, regulation; low [92]
(DNA only) tracking; quick
KRAS) abundance in
turnaround
early-stage disease
Phenotypic and genotypic Low frequency; difficult
Possible (but profiling of Can be cultured; to isolate and purify; not
2 CTCs Whole blood rarely used for resistance-related provides both RNA and ideal for circRNA [86]
circRNA) protein information
pathways analysis
Isolation and
Reflects tumor-secreted Protect circRNAs from characterization
Plasma, serum, Highly enriched resistance-related degradation;
3 Exosomes methods not yet [91]
saliva, urine, CSF for circRNAs molecules and tumor-specific; suitable standardized; requires
intercellular signals for dynamic tracking
optimization
Tracks therapy-related CircRNAs are a minor
Includes changes in gene Minimally invasive; fraction of total cfRNA;
4 cfRNA Plasma/serum circRNAs (after expression and resistance suitable for repeated requires enrichment and [93]
enrichment) monitoring
regulation sensitive detection
Exosome fraction Direct and Dynamic monitoring of High specificity and Requires high-quality
Exosomal
5 from stable source of resistance; potential stability; resistant to isolation; functional [94]
circRNAs
plasma/serum circRNAs predictive biomarker RNases validation needed
circRNAs: Circular RNAs; ctDNA: circulating tumor DNA; EGFR: epidermal growth factor receptor; KRAS: kirsten rat sarcoma viral oncogene
homolog; CTCs: circulating tumor cells; CSF: cerebrospinal fluid; cfRNA: cell-free RNA.
37