Page 43 - Read Online
P. 43
Page 8 of 20 Singh et al. Cancer Drug Resist. 2025;8:56
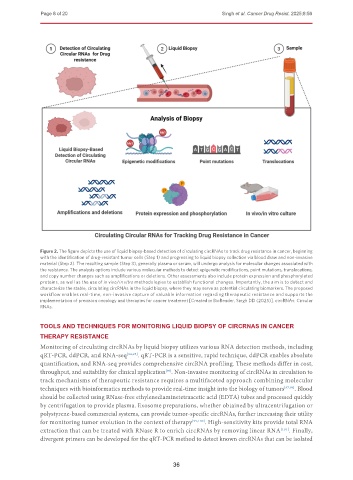
Figure 2. The figure depicts the use of liquid biopsy-based detection of circulating circRNAs to track drug resistance in cancer, beginning
with the identification of drug-resistant tumor cells (Step 1) and progressing to liquid biopsy collection via blood draw and non-invasive
material (Step 2). The resulting sample (Step 3), generally plasma or serum, will undergo analysis for molecular changes associated with
the resistance. The analysis options include various molecular methods to detect epigenetic modifications, point mutations, translocations,
and copy number changes such as amplifications or deletions. Other assessments also include protein expression and phosphorylated
proteins, as well as the use of in vivo/in vitro methodologies to establish functional changes. Importantly, the aim is to detect and
characterize the stable, circulating circRNAs in the liquid biopsy, where they may serve as potential circulating biomarkers. The proposed
workflow enables real-time, non-invasive capture of valuable information regarding therapeutic resistance and supports the
implementation of precision oncology and therapies for cancer treatment [Created in BioRender. Singh DD (2025)]. circRNAs: Circular
RNAs.
TOOLS AND TECHNIQUES FOR MONITORING LIQUID BIOPSY OF CIRCRNAS IN CANCER
THERAPY RESISTANCE
Monitoring of circulating circRNAs by liquid biopsy utilizes various RNA detection methods, including
qRT-PCR, ddPCR, and RNA-seq [94,95] . qRT-PCR is a sensitive, rapid technique, ddPCR enables absolute
quantification, and RNA-seq provides comprehensive circRNA profiling. These methods differ in cost,
throughput, and suitability for clinical application . Non-invasive monitoring of circRNAs in circulation to
[96]
track mechanisms of therapeutic resistance requires a multifaceted approach combining molecular
techniques with bioinformatics methods to provide real-time insight into the biology of tumors [97,98] . Blood
should be collected using RNase-free ethylenediaminetetraacetic acid (EDTA) tubes and processed quickly
by centrifugation to provide plasma. Exosome preparations, whether obtained by ultracentrifugation or
polystyrene-based commercial systems, can provide tumor-specific circRNAs, further increasing their utility
for monitoring tumor evolution in the context of therapy [99,100] . High-sensitivity kits provide total RNA
extraction that can be treated with RNase R to enrich circRNAs by removing linear RNA [101] . Finally,
divergent primers can be developed for the qRT-PCR method to detect known circRNAs that can be isolated
36