Page 47 - Read Online
P. 47
Page 12 of 20 Singh et al. Cancer Drug Resist. 2025;8:56
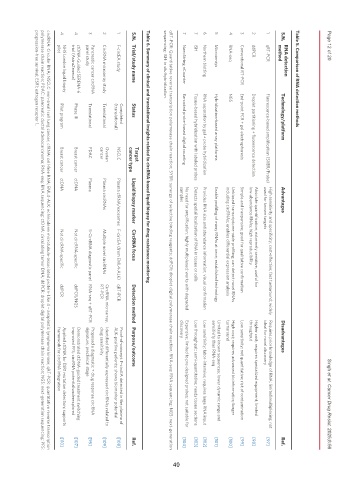
Table 5. Comparison of RNA detection methods
RNA detection
S.N. Technology/platform Advantages Disadvantages Ref.
method
High sensitivity and specificity; cost-effective; fast turnaround; widely Requires prior knowledge of RNA; limited multiplexing; not
1 qRT-PCR Fluorescence-based amplification (SYBR/Probe) [97]
used for known targets ideal for novel discovery
Absolute quantification; extremely sensitive; useful for Higher cost; requires specialized equipment; limited
2 ddPCR Droplet partitioning + fluorescence detection [98]
low-abundance RNAs; high reproducibility throughput
3 Conventional RT-PCR End-point PCR + gel electrophoresis Simple and inexpensive; good for qualitative confirmation Low sensitivity; not quantitative; risk of contamination [99]
Unbiased transcriptome-wide profiling; can detect novel RNAs High cost; requires advanced bioinformatics; longer
4 RNA-seq NGS [100]
including circRNAs; enables differential expression analysis turnaround
Limited to known sequences; lower dynamic range and
5 Microarrays Hybridization-based array platforms Enable profiling of many RNAs at once; established technology [101]
sensitivity than RNA-seq
6 Northern blotting RNA separation by gel + probe hybridization Provides RNA size and abundance information; visual confirmation Low sensitivity; labor-intensive; requires large RNA input [102]
7 ISH Tissue-based hybridization with labeled probes Detects spatial localization of RNA in tissue or cells Low throughput; semi-quantitative; needs tissue sections [103]
No need for amplification; highly multiplexed; works with degraded Expensive; limited to pre-designed probes; not suitable for
7 NanoString nCounter Barcoded probe-based digital counting [104]
samples discovery
40
qRT-PCR: Quantitative reverse transcription polymerase chain reaction; SYBR: synergy of selective binding reagents; ddPCR: droplet digital polymerase chain reaction; RNA-seq: RNA sequencing; NGS: next-generation
sequencing; ISH: in situ hybridization.
Table 6. Summary of clinical and translational insights related to circRNA-based liquid biopsy for drug resistance monitoring
Target
S.N. Trial/study name Status Liquid biopsy marker CircRNA focus Detection method Purpose/outcome Ref.
cancer type
Completed Proof-of-concept: F-circEA detected in the plasma of
1 F-circEA study NSCLC Plasma cfRNA/exosomes F-circEA (from EML4-ALK) qRT-PCR [108]
(translational) ALK-positive patients; shows biomarker potential
Ovarian CircRNA microarray, Identified differentially expressed circRNAs related to
2 CircRNA microarray study Translational Plasma circRNAs Multiple novel circRNAs [109]
cancer RT-PCR drug sensitivity
Pancreatic cancer circRNA Proposed a diagnostic + drug response circRNA
3 Translational PDAC Plasma 5-CircRNA diagnostic panel RNA-seq + qRT-PCR [95]
panel study signature; preclinical stage
ctDNA-Guided SERENA-6 Demonstrated ctDNA-guided treatment switching
4 Phase III Breast cancer ctDNA Not circRNA-specific ddPCR/NGS [107]
trial (AstraZeneca) improved PFS; circRNA potential underexplored
NHS London liquid biopsy Applied ctDNA for ESR1 mutation detection; supports
4 Pilot program Breast cancer ctDNA Not circRNA-specific ddPCR [110]
pilot framework for circRNA integration
circRNA: Circular RNA; NSCLC: non-small cell lung cancer; cfRNA: cell-free RNA; EML4-ALK: echinoderm microtubule-associated protein-Like 4 – anaplastic lymphoma kinase; qRT-PCR: quantitative reverse transcription
polymerase chain reaction; PDAC: pancreatic ductal adenocarcinoma; RNA-seq: RNA sequencing; ctDNA: circulating tumor DNA; ddPCR: droplet digital polymerase chain reaction; NGS: next-generation sequencing; PFS:
progression-free survival; ESR1: estrogen receptor 1.