Page 104 - Read Online
P. 104
Page 10 of 26 Li et al. Cancer Drug Resist. 2025;8:31
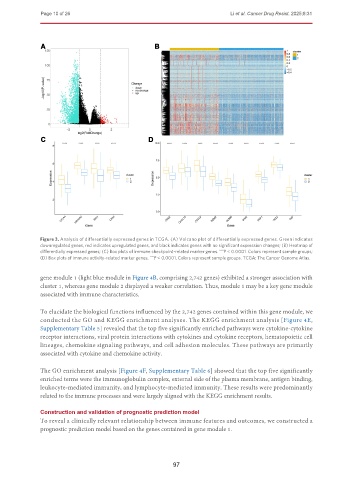
Figure 3. Analysis of differentially expressed genes in TCGA. (A) Volcano plot of differentially expressed genes. Green indicates
downregulated genes, red indicates upregulated genes, and black indicates genes with no significant expression changes; (B) Heatmap of
differentially expressed genes; (C) Box plots of immune checkpoint-related marker genes. P < 0.0001. Colors represent sample groups;
****
(D) Box plots of immune activity-related marker genes. P < 0.0001. Colors represent sample groups. TCGA: The Cancer Genome Atlas.
****
gene module 1 (light blue module in Figure 4B, comprising 2,742 genes) exhibited a stronger association with
cluster 1, whereas gene module 2 displayed a weaker correlation. Thus, module 1 may be a key gene module
associated with immune characteristics.
To elucidate the biological functions influenced by the 2,742 genes contained within this gene module, we
conducted the GO and KEGG enrichment analyses. The KEGG enrichment analysis [Figure 4E,
Supplementary Table 5] revealed that the top five significantly enriched pathways were cytokine-cytokine
receptor interactions, viral protein interactions with cytokines and cytokine receptors, hematopoietic cell
lineages, chemokine signaling pathways, and cell adhesion molecules. These pathways are primarily
associated with cytokine and chemokine activity.
The GO enrichment analysis [Figure 4F, Supplementary Table 6] showed that the top five significantly
enriched terms were the immunoglobulin complex, external side of the plasma membrane, antigen binding,
leukocyte-mediated immunity, and lymphocyte-mediated immunity. These results were predominantly
related to the immune processes and were largely aligned with the KEGG enrichment results.
Construction and validation of prognostic prediction model
To reveal a clinically relevant relationship between immune features and outcomes, we constructed a
prognostic prediction model based on the genes contained in gene module 1.
97