Page 40 - Read Online
P. 40
Page 10 Qu et al. J Transl Genet Genom 2023;7:3-16 https://dx.doi.org/10.20517/jtgg.2022.16
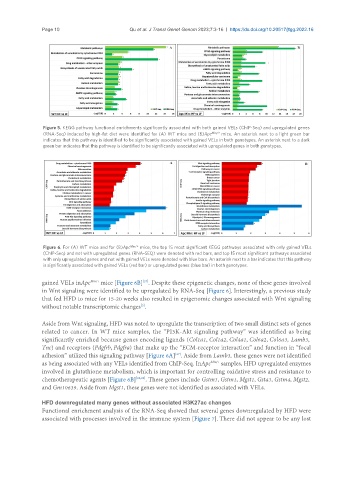
Figure 5. KEGG pathway functional enrichments significantly associated with both gained VELs (ChIP-Seq) and upregulated genes
(RNA-Seq) induced by high-fat diet were identified for (A) WT mice and (B)Apc Min/+ mice. An asterisk next to a light green bar
indicates that this pathway is identified to be significantly associated with gained VELs in both genotypes. An asterisk next to a dark
green bar indicates that this pathway is identified to be significantly associated with upregulated genes in both genotypes.
Figure 6. For (A) WT mice and for (B)Apc Min/+ mice, the top 15 most significant KEGG pathways associated with only gained VELs
(ChIP-Seq) and not with upregulated genes (RNA-SEQ) were denoted with red bars, and top 15 most significant pathways associated
with only upregulated genes and not with gained VELs were denoted with blue bars. An asterisk next to a bar indicates that this pathway
is significantly associated with gained VELs (red bar) or upregulated genes (blue bar) in both genotypes.
gained VELs inApc Min/+ mice [Figure 6B] . Despite these epigenetic changes, none of these genes involved
[27]
in Wnt signaling were identified to be upregulated by RNA-Seq [Figure 6]. Interestingly, a previous study
that fed HFD to mice for 15-20 weeks also resulted in epigenomic changes associated with Wnt signaling
without notable transcriptomic changes .
[5]
Aside from Wnt signaling, HFD was noted to upregulate the transcription of two small distinct sets of genes
related to cancer. In WT mice samples, the “PI3K-Akt signaling pathway” was identified as being
significantly enriched because genes encoding ligands (Col1a1, Col1a2, Col4a1, Col6a2, Col6a3, Lamb3,
Tnc) and receptors (Pdgfrb, Pdgfra) that make up the “ECM-receptor interaction” and function in “focal
adhesion” utilized this signaling pathway [Figure 6A] . Aside from Lamb3, these genes were not identified
[27]
as being associated with any VELs identified from ChIP-Seq. InApc Min/+ samples, HFD upregulated enzymes
involved in glutathione metabolism, which is important for controlling oxidative stress and resistance to
chemotherapeutic agents [Figure 6B] [28,29] . These genes include Gstm3, Gstm1, Mgst1, Gsta3, Gstm4, Mgst2,
and Gm10639. Aside from Mgst1, these genes were not identified as associated with VELs.
HFD downregulated many genes without associated H3K27ac changes
Functional enrichment analysis of the RNA-Seq showed that several genes downregulated by HFD were
associated with processes involved in the immune system [Figure 7]. There did not appear to be any lost