Page 38 - Read Online
P. 38
Page 8 Qu et al. J Transl Genet Genom 2023;7:3-16 https://dx.doi.org/10.20517/jtgg.2022.16
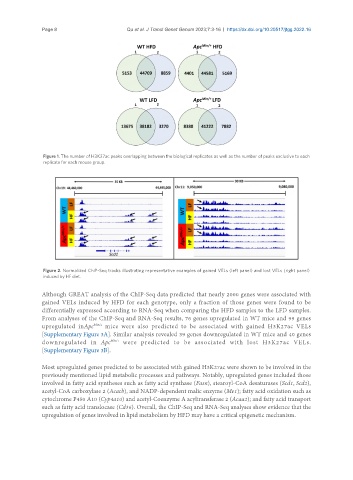
Figure 1. The number of H3K27ac peaks overlapping between the biological replicates as well as the number of peaks exclusive to each
replicate for each mouse group.
Figure 2. Normalized ChiP-Seq tracks illustrating representative examples of gained VELs (left panel) and lost VELs (right panel)
induced by HF diet.
Although GREAT analysis of the ChIP-Seq data predicted that nearly 2000 genes were associated with
gained VELs induced by HFD for each genotype, only a fraction of those genes were found to be
differentially expressed according to RNA-Seq when comparing the HFD samples to the LFD samples.
From analyses of the ChIP-Seq and RNA-Seq results, 76 genes upregulated in WT mice and 99 genes
upregulated inApc Min/+ mice were also predicted to be associated with gained H3K27ac VELs
[Supplementary Figure 3A]. Similar analysis revealed 39 genes downregulated in WT mice and 40 genes
d o w n r e g u l a t e d i n Apc Min/+ w e r e p r e d i c t e d t o b e a s s o c i a t e d w i t h l o s t H 3 K 2 7 a c V E L s .
[Supplementary Figure 3B].
Most upregulated genes predicted to be associated with gained H3K27ac were shown to be involved in the
previously mentioned lipid metabolic processes and pathways. Notably, upregulated genes included those
involved in fatty acid syntheses such as fatty acid synthase (Fasn), stearoyl-CoA desaturases (Scd1, Scd2),
acetyl-CoA carboxylase 2 (Acacb), and NADP-dependent malic enzyme (Me1); fatty acid oxidation such as
cytochrome P450 A10 (Cyp4a10) and acetyl-Coenzyme A acyltransferase 2 (Acaa2); and fatty acid transport
such as fatty acid translocase (Cd36). Overall, the ChIP-Seq and RNA-Seq analyses show evidence that the
upregulation of genes involved in lipid metabolism by HFD may have a critical epigenetic mechanism.