Page 39 - Read Online
P. 39
Qu et al. J Transl Genet Genom 2023;7:3-16 https://dx.doi.org/10.20517/jtgg.2022.16 Page 9
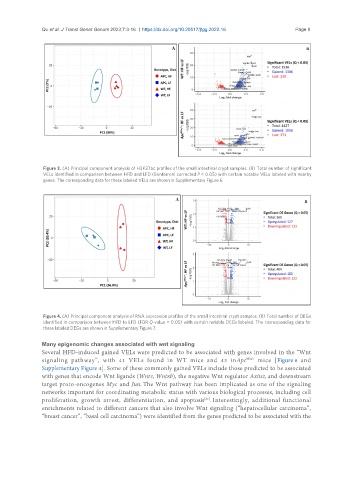
Figure 3. (A) Principal component analysis of H3K27ac profiles of the small intestinal crypt samples. (B) Total number of significant
VELs identified in comparison between HFD and LFD (Bonferroni corrected P < 0.05) with certain notable VELs labeled with nearby
genes. The corresponding data for these labeled VELs are shown in Supplementary Figure 6.
Figure 4. (A) Principal component analysis of RNA expression profiles of the small intestinal crypt samples. (B) Total number of DEGs
identified in comparison between HFD to LFD (FDR Q-value < 0.05) with certain notable DEGs labeled. The corresponding data for
these labeled DEGs are shown in Supplementary Figure 7.
Many epigenomic changes associated with wnt signaling
Several HFD-induced gained VELs were predicted to be associated with genes involved in the “Wnt
signaling pathway”, with 41 VELs found in WT mice and 43 inApc Min/+ mice [Figure 6 and
Supplementary Figure 4]. Some of these commonly gained VELs include those predicted to be associated
with genes that encode Wnt ligands (Wnt3, Wnt8b), the negative Wnt regulator Axin2, and downstream
target proto-oncogenes Myc and Jun. The Wnt pathway has been implicated as one of the signaling
networks important for coordinating metabolic status with various biological processes, including cell
[26]
proliferation, growth arrest, differentiation, and apoptosis . Interestingly, additional functional
enrichments related to different cancers that also involve Wnt signaling (“hepatocellular carcinoma”,
“breast cancer”, “basal cell carcinoma”) were identified from the genes predicted to be associated with the