Page 10 - Read Online
P. 10
Page 131 Wang et al. J Transl Genet Genom 2023;7:126-40 https://dx.doi.org/10.20517/jtgg.2023.07
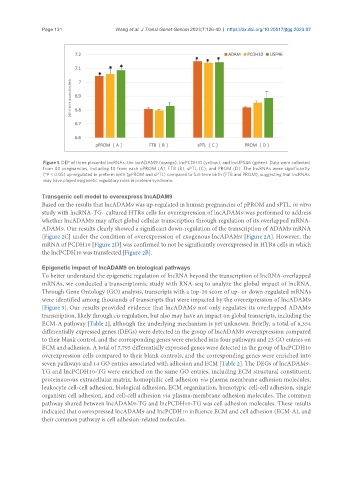
Figure 1. DEP of three placental IncRNAs, the IncADAM9 (orange), IncPCDH10 (yellow), and IncUPS46 (green). Data were collected
from 40 pregnancies, including 10 from each pPROM (A), FTB (B), sPTL (C), and PROM (D). The IncRNAs were significantly
(*P < 0.05) up-regulated in preterm birth (pPROM and sPTL) compared to full-term birth (FTB and PROM), suggesting that IncRNAs
may have played epigenetic regulatory roles in preterm syndrome.
Transgenic cell model to overexpress lncADAM9
Based on the results that lncADAM9 was up-regulated in human pregnancies of pPROM and sPTL, in vitro
study with lncRNA-TG- cultured HTR8 cells for overexpression of lncADAM9 was performed to address
whether lncADAM9 may affect global cellular transcription through regulation of its overlapped mRNA-
ADAM9. Our results clearly showed a significant down-regulation of the transcription of ADAM9 mRNA
[Figure 2C] under the condition of overexpression of exogenous lncADAM9 [Figure 2A]. However, the
mRNA of PCDH10 [Figure 2D] was confirmed to not be significantly overexpressed in HTR8 cells in which
the lncPCDH10 was transfected [Figure 2B].
Epigenetic impact of lncADAM9 on biological pathways
To better understand the epigenetic regulation of lncRNA beyond the transcription of lncRNA-overlapped
mRNAs, we conducted a transcriptomic study with RNA-seq to analyze the global impact of lncRNA.
Through Gene Ontology (GO) analysis, transcripts with a top-20 score of up- or down-regulated mRNAs
were identified among thousands of transcripts that were impacted by the overexpression of lncADAM9
[Figure 3]. Our results provided evidence that lncADAM9 not only regulates its overlapped ADAM9
transcription, likely through cis regulation, but also may have an impact on global transcripts, including the
ECM-A pathway [Table 2], although the underlying mechanism is yet unknown. Briefly, a total of 8,354
differentially expressed genes (DEGs) were detected in the group of lncADAM9 overexpression compared
to their blank control, and the corresponding genes were enriched into four pathways and 23 GO entries on
ECM and adhesion. A total of 7,795 differentially expressed genes were detected in the group of lncPCDH10
overexpression cells compared to their blank controls, and the corresponding genes were enriched into
seven pathways and 14 GO entries associated with adhesion and ECM [Table 2]. The DEGs of lncADAM9-
TG and lncPCDH10-TG were enriched on the same GO entries, including ECM structural constituent,
proteinaceous extracellular matrix, homophilic cell adhesion via plasma membrane adhesion molecules,
leukocyte cell-cell adhesion, biological adhesion, ECM organization, homotypic cell-cell adhesion, single
organism cell adhesion, and cell-cell adhesion via plasma-membrane adhesion molecules. The common
pathway shared between lncADAM9-TG and lncPCDH10-TG was cell adhesion molecules. These results
indicated that overexpressed lncADAM9 and lncPCDH10 influence ECM and cell adhesion (ECM-A), and
their common pathway is cell adhesion-related molecules.