Page 22 - Read Online
P. 22
Bergara-Muguruza et al. J Transl Genet Genom 2023;7:213-229 https://dx.doi.org/10.20517/jtgg.2023.14 Page 215
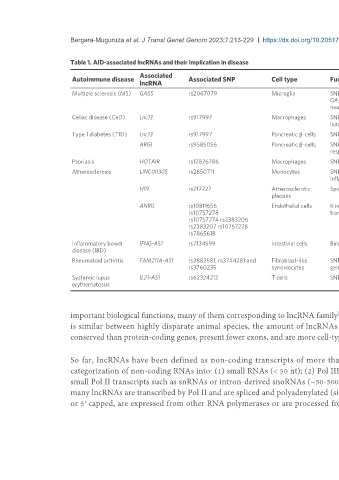
Table 1. AID-associated lncRNAs and their implication in disease
Associated
Autoimmune disease Associated SNP Cell type Function Refs.
lncRNA
Multiple sclerosis (MS) GAS5 rs2067079 Microglia SNP located in promoter/enhancer and predicted structure alteration [47-49,51,52]
GAS5 competes with miR-137, which releases its target Notch1, resulting in a decrease in
neuronal survival
Celiac disease (CeD) Lnc13 rs917997 Macrophages SNP disrupts the structure of the lncRNA, decreasing the interaction with hnRNDP and thus [13,61]
leading to the expression of disease-related proinflammatory genes
Type 1 diabetes (T1D) Lnc13 rs917997 Pancreatic β-cells SNP promotes interaction with PCBP2 and STAT1 mRNA affecting stability [69]
ARGI rs9585056 Pancreatic β-cells SNP is predicted to change the secondary structure of ARGI and it exacerbates type I IFN [71]
response
Psoriasis HOTAIR rs12826786 Macrophages SNP increases HOTAIR expression, which may induce NFkB activation [75-79]
Atherosclerosis LINC00305 rs2850711 Monocytes SNP increases its expression. LINC00305 modulates NF-κB and promotes monocyte [80]
inflammation
H19 rs217727 Atherosclerotic Sponges the miRNAs from the let-7 family [34,81,82]
plaques
ANRIL rs10811656 Endothelial cells It recruits chromatin modifiers to inhibit gene expression in cis and binds to several factors to [84-86]
rs10757278 trans-regulate some genes
rs10757274 rs2383206
rs2383207 rs10757278
rs7865618
Inflammatory bowel IFNG-AS1 rs7134599 Intestinal cells Binds to a histone methylation complex and this methylation activates IFNG transcription [87-91]
disease (IBD)
Rheumatoid arthritis FAM211A-AS1 rs2882581, rs3744281 and Fibroblast-like SNPs seem to locate in regulatory elements influencing lncRNA transcription and thus nearby [95,99]
rs3760235 synoviocytes genes
Systemic lupus IL21-AS1 rs62324212 T cells SNP located in enhancer regions, which may affect the expression of the lncRNA [100]
erythematosus
important biological functions, many of them corresponding to lncRNA family , which opened a new avenue of research. While protein-coding gene number
[18]
is similar between highly disparate animal species, the amount of lncRNAs increases with evolutionary complexity. Moreover, these molecules are less
conserved than protein-coding genes, present fewer exons, and are more cell-type specifically and less abundantly expressed than coding genes [15,19] .
So far, lncRNAs have been defined as non-coding transcripts of more than 200 nt, but recent consensus statement have suggested a more precise
[19]
categorization of non-coding RNAs into: (1) small RNAs (< 50 nt); (2) Pol III transcripts (i.e., tRNAs, 5S rRNA, 7SK, 7SL, and Alu, vault and Y RNAs) and
small Pol II transcripts such as snRNAs or intron-derived snoRNAs (~50-500 nt); and (3) lncRNAs (> 500 nt), which are mostly generated by Pol II. While
many lncRNAs are transcribed by Pol II and are spliced and polyadenylated (similarly to mRNAs), there are many other lncRNAs that are not polyadenylated
or 5′ capped, are expressed from other RNA polymerases or are processed from introns and repetitive elements. Moreover, regarding their location in the