Page 52 - Read Online
P. 52
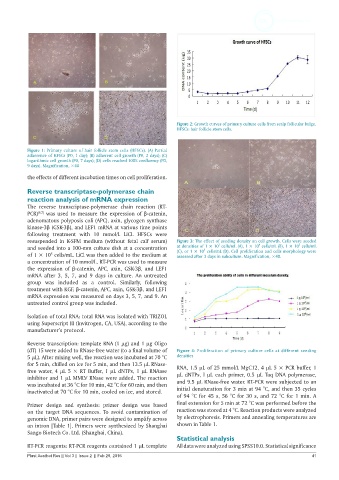
Figure 2: Growth curves of primary culture cells from scalp follicular bulge.
HFSCs: hair follicle stem cells.
Figure 1: Primary culture of hair follicle stem cells (HFSCs). (A) Partial
adherence of HFSCs (P0, 1 day); (B) adherent cell growth (P0, 3 days); (C)
logarithmic cell growth (P0, 7 days); (D) cells reached 100% confluency (P0,
9 days). Magnification, ×40
the effects of different incubation times on cell proliferation.
Reverse transcriptase-polymerase chain
reaction analysis of mRNA expression
The reverse transcriptase-polymerase chain reaction (RT-
PCR) [6,7] was used to measure the expression of β-catenin,
adenomatous polyposis coli (APC), axin, glycogen synthase
kinase-3β (GSK-3β), and LEF1 mRNA at various time points
following treatment with 10 mmol/L LiCl. HFSCs were
resuspended in K-SFM medium (without fetal calf serum) Figure 3: The effect of seeding density on cell growth. Cells were seeded
5
4
3
and seeded into a 100-mm culture dish at a concentration at densities of 1 × 10 cells/mL (A), 1 × 10 cells/mL (B), 1 × 10 cells/mL
6
of 1 × 10 cells/mL. LiCl was then added to the medium at (C), or 1 × 10 cells/mL (D). Cell proliferation and cells morphology were
5
assessed after 3 days in subculture. Magnification, ×40.
a concentration of 10 mmol/L. RT-PCR was used to measure
the expression of β-catenin, APC, axin, GSK-3β, and LEF1
mRNA after 3, 5, 7, and 9 days in culture. An untreated
group was included as a control. Similarly, following
treatment with KGF, β-catenin, APC, axin, GSK-3β, and LEF1
mRNA expression was measured on days 3, 5, 7, and 9. An
untreated control group was included.
Isolation of total RNA: total RNA was isolated with TRIZOL
using Superscript III (Invitrogen, CA, USA), according to the
manufacturer’s protocol.
Reverse transcription: template RNA (1 µg) and 1 µg Oligo
(dT) 15 were added to RNase-free water (to a final volume of Figure 4: Proliferation of primary culture cells at different seeding
5 µL). After mixing well, the reaction was incubated at 70 °C densities
for 5 min, chilled on ice for 5 min, and then 13.5 µL RNase- RNA, 1.5 µL of 25 mmol/L MgC12, 4 µL 5 × PCR buffer, 1
free water, 4 µL 5 × RT Buffer, 1 µL dNTPs, 1 µL RNAase µL dNTPs, l µL each primer, 0.5 µL Taq DNA polymerase,
inhibitor and 1 µL MMLV RNase were added. The reaction
was incubated at 36 °C for 10 min, 42 °C for 60 min, and then and 9.5 µL RNase-free water. RT-PCR were subjected to an
inactivated at 70 °C for 10 min, cooled on ice, and stored. initial denaturation for 3 min at 94 °C, and then 35 cycles
of 94 °C for 45 s, 56 °C for 30 s, and 72 °C for 1 min. A
Primer design and synthesis: primer design was based final extension for 5 min at 72 °C was performed before the
on the target DNA sequences. To avoid contamination of reaction was stored at 4 °C. Reaction products were analyzed
genomic DNA, primer pairs were designed to amplify across by electrophoresis. Primers and annealing temperatures are
an intron [Table 1]. Primers were synthesized by Shanghai shown in Table 1.
Sango Biotech Co. Ltd. (Shanghai, China).
Statistical analysis
RT-PCR reagents: RT-PCR reagents contained 1 µL template All data were analyzed using SPSS10.0. Statistical significance
Plast Aesthet Res || Vol 3 || Issue 2 || Feb 29, 2016 41