Page 52 - Read Online
P. 52
Shad. J Transl Genet Genom 2023;7:141-65 https://dx.doi.org/10.20517/jtgg.2023.11 Page 145
[48]
2007 Dutch -HLA-DRB5*0201-HLA- 0.004
DRB4*000 0.004
-HLA-Cw7-B18 NA NA NA NA NA NA NA 0.005
-HLA-DRB5*000 0.018
-HLA-Cw7-B39-DRB5*000
-HLA-Cw7-B44-DRB5*000
Van der Weide et al., -ABCB C3435T 18 13 101 140 91% 15% 3.13 0.05
[49]
2017 (rs1045642) 14 17 64 177 96% 18% 5.53 0.004
-NQO2 G1541A
Athanasiou et al., Cohort 1: US, Russia, and Cohort1 I: NA NA NA NA NA NA NA
[50]
2011 South Africa
DRD1 < 0.05
CSF2RB < 0.05
NTSR1 < 0.05
Cohort II: German -HLA-DQB1 6672G>C 8 1 24 52 98% 25.0% 17.33 0.0015
Caucasians, non-Jewish Cohort II: -HLA-DQB1 9 1 38 71 99% 19% 16.82 0.00097
6672G>C
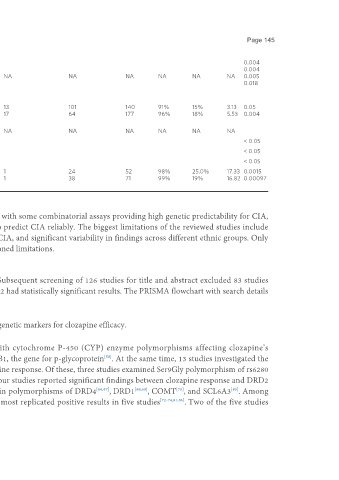
Although some of the findings from the reviewed studies have been replicated with some combinatorial assays providing high genetic predictability for CIA,
more research is warranted to confirm the clinical utility of genetic testing to predict CIA reliably. The biggest limitations of the reviewed studies include
inadequate sample sizes, failing to assess relatively rare genetic predictors for CIA, and significant variability in findings across different ethnic groups. Only
genome-wide assays in significantly large sample studies can address the mentioned limitations.
Antipsychotic response with clozapine
In terms of clozapine efficacy, 126 studies were potentially eligible initially. Subsequent screening of 126 studies for title and abstract excluded 83 studies
leaving 69, of which 31 were ineligible, six were duplicates, and the remaining 32 had statistically significant results. The PRISMA flowchart with search details
is shown in Figure 2.
As can be seen from Table 2 below, reviewed studies examined the PK and PD genetic markers for clozapine efficacy.
Significant changes in clozapine response were reported in four studies with cytochrome P-450 (CYP) enzyme polymorphisms affecting clozapine’s
[57]
[58]
metabolism targeting CYP1A2 [54-56] , and CYP2C19 , and one study with ABCB1, the gene for p-glycoprotein . At the same time, 13 studies investigated the
effects of various genetic polymorphisms in the dopamine system on the clozapine response. Of these, three studies examined Ser9Gly polymorphism of rs6280
in the DRD3 gene with significant positive effects on clozapine response [59-61] . Four studies reported significant findings between clozapine response and DRD2
[70]
polymorphisms [62-65] . The remaining six studies examined clozapine response in polymorphisms of DRD4 [66,67] , DRD1 [68,69] , COMT , and SCL6A3 . Among
[85]
the serotonergic genes, the HTR2A gene for 5HT2A receptors produced the most replicated positive results in five studies [72-74,81,86] . Two of the five studies